Note
Go to the end to download the full example code.
Load Project
Load and visualize an OMF project file
# sphinx_gallery_thumbnail_number = 3
import pyvista as pv
import omfvista
Load the project into an pyvista.MultiBlock dataset
project = omfvista.load_project("../assets/test_file.omf")
print(project)
MultiBlock (0x7ffa3e420700)
N Blocks: 9
X Bounds: 4.439e+05, 4.471e+05
Y Bounds: 4.919e+05, 4.951e+05
Z Bounds: 2.330e+03, 3.556e+03
Once the data is loaded as a pyvista.MultiBlock dataset from
omfvista, then that object can be directly used for interactive 3D
visualization from pyvista:
project.plot()
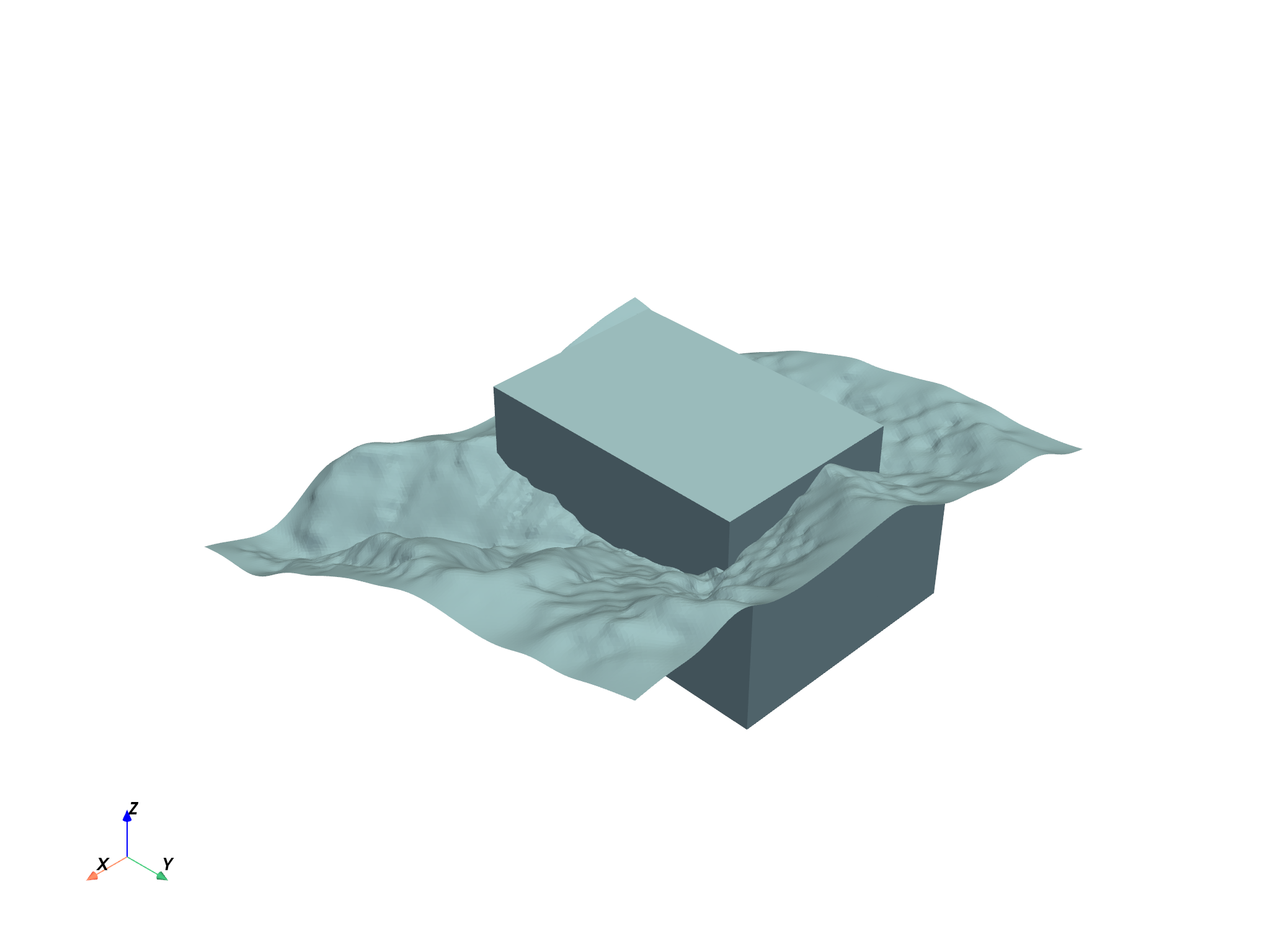

Or an interactive scene can be created and manipulated to create a compelling figure directly in a Jupyter notebook. First, grab the elements from the project:
# Grab a few elements of interest and plot em up!
vol = project["Block Model"]
assay = project["wolfpass_WP_assay"]
topo = project["Topography"]
dacite = project["Dacite"]
assay.set_active_scalars("DENSITY")
p = pv.Plotter()
p.add_mesh(assay.tube(radius=3))
p.add_mesh(topo, opacity=0.5)
p.show()
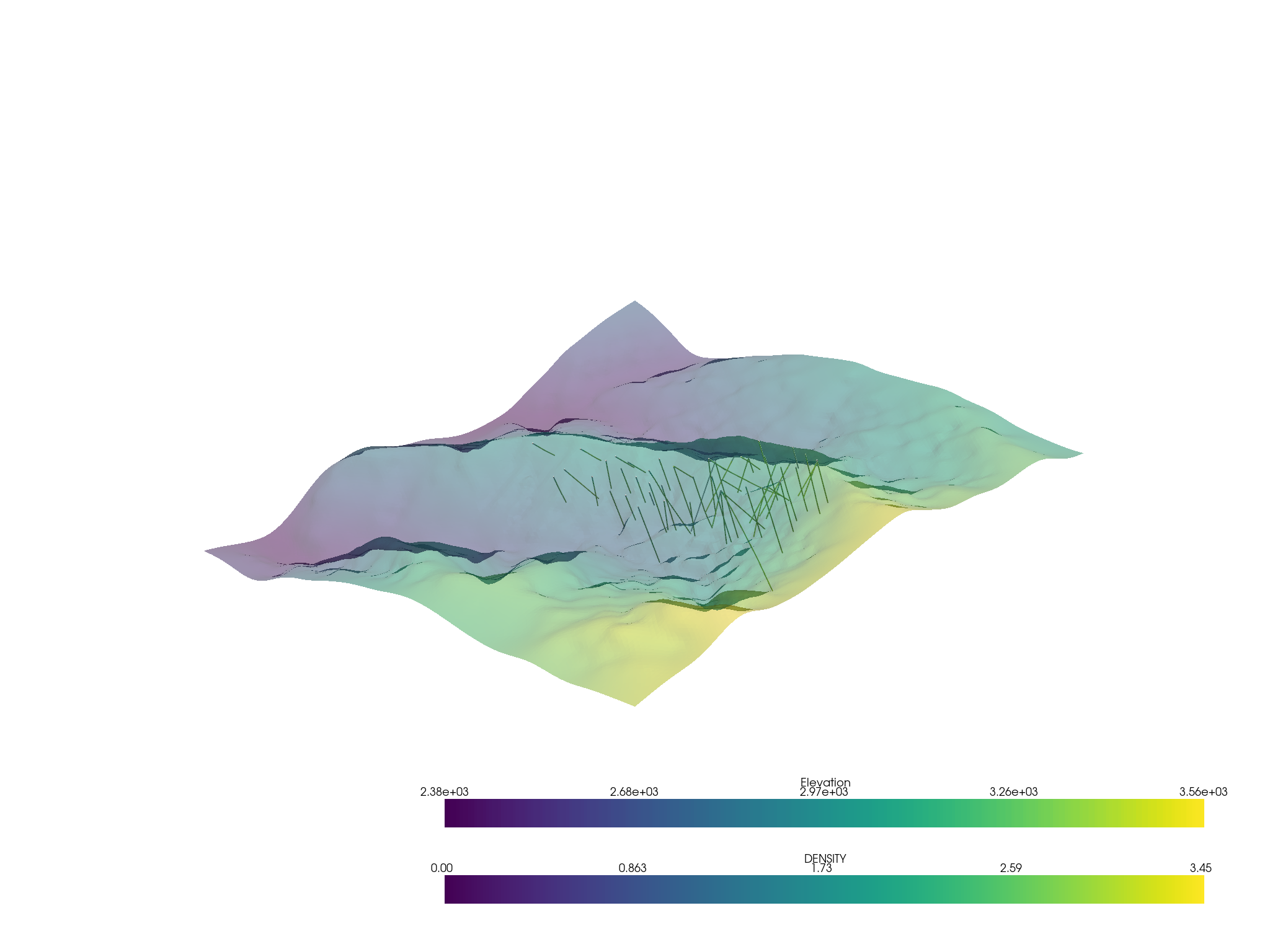
Then apply a filtering tool from pyvista to the volumetric data:
# Threshold the volumetric data
thresh_vol = vol.threshold([1.09, 4.20])
print(thresh_vol)
UnstructuredGrid (0x7ffa3c0ccd00)
N Cells: 92525
N Points: 107807
X Bounds: 4.447e+05, 4.457e+05
Y Bounds: 4.929e+05, 4.942e+05
Z Bounds: 2.330e+03, 3.110e+03
N Arrays: 1
Then you can put it all in one environment!
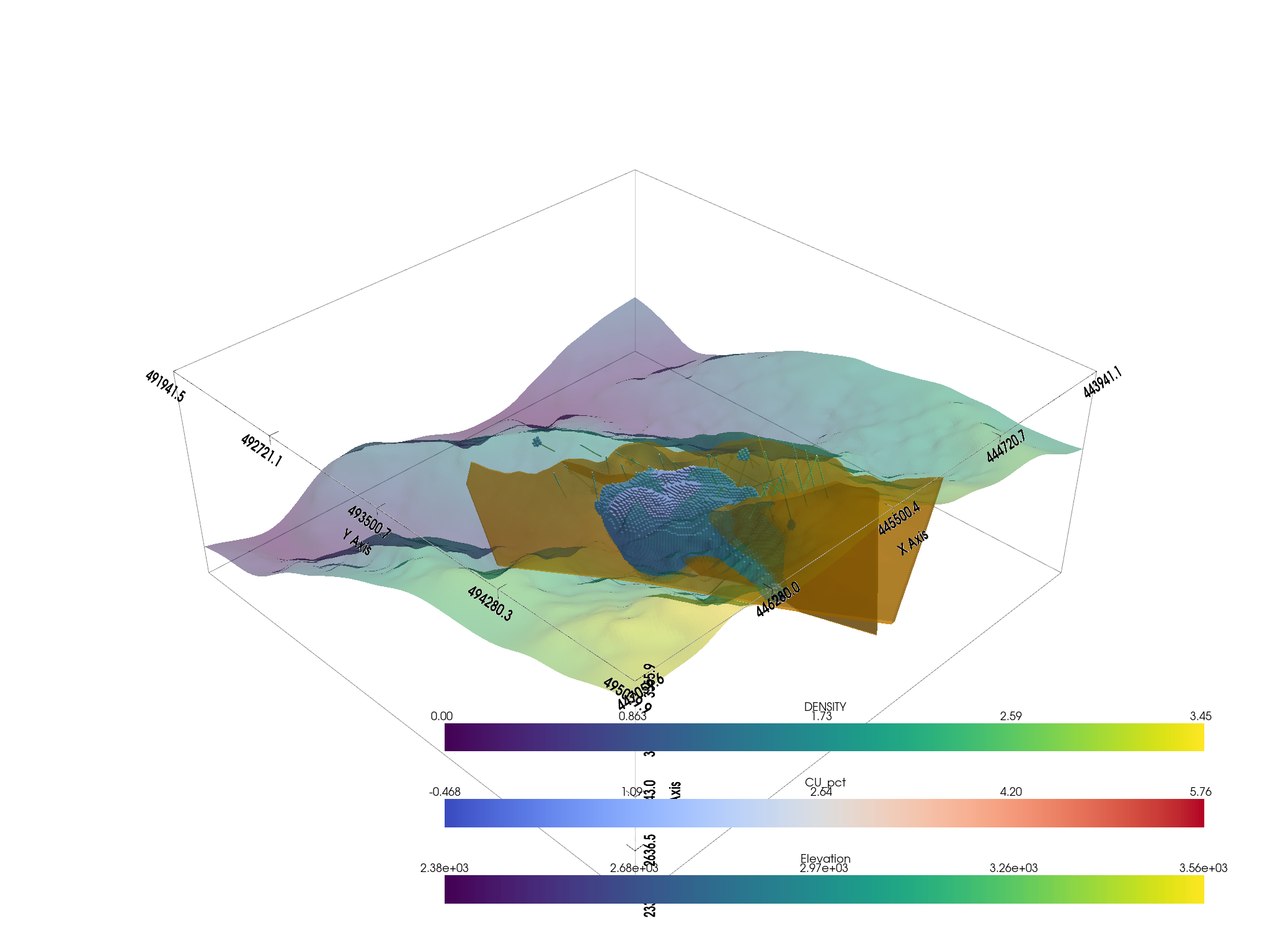
# Create a plotting window
p = pv.Plotter()
# Add the bounds axis
p.show_bounds()
p.add_bounding_box()
# Add our datasets
p.add_mesh(topo, opacity=0.5)
p.add_mesh(
dacite,
color="orange",
opacity=0.6,
)
p.add_mesh(thresh_vol, cmap="coolwarm", clim=vol.get_data_range())
# Add the assay logs: use a tube filter that varius the radius by an attribute
p.add_mesh(assay.tube(radius=3), cmap="viridis")
p.show()
Total running time of the script: (0 minutes 7.125 seconds)